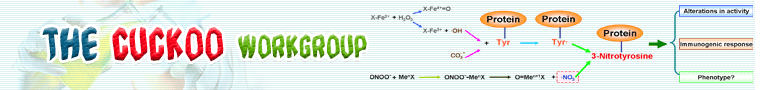※ GPS-YNO2 INTRODUCTION:
News: On Nov. 24, 2015, GSP-YNO2 was listed in ![]() , a tiny client app for the largest software library, and one of the world's 500 most visited websites with over 700,000 unique visitors every day!
, a tiny client app for the largest software library, and one of the world's 500 most visited websites with over 700,000 unique visitors every day!
This paper is the TOP TEN ACCESSED ARTICLES in August, 2011
from the Molecular Biosystems
Early after the discovery of the freely-diffusible signaling molecule and second messenger nitric oxide (NO) and its function in signal transduction within the vasculature and nervous system, it became obvious that NO also take part in cytotoxic cytotoxicity when abundantly produced by iNOS, eNOS or nNOS (Ignarro, L.J. et al., 1990). When NO meets molecular oxygen in hydrophobic environments where these species concentrate, the nitrogen dioxide could be formed, then reactive nitrogen species are derived which leave nitration as the molecular footprints by reactions with proteins. Tyrosine nitration is a kind of covalent post-translational modifications which adds a nitro (-NO2) group onto either one of the two ortho carbons of the aromatic ring of tyrosine resides. Originally, protein tyrosine nitration was considered to take place only in vitro mediated by reactive nitrating agent such as tetranitromethane (Sokolovsky, M. et al., 1992) and peroxynitrite (Beckman, J.S. et al., 1992; Ischiropoulos, H. et al., 1992). Later, seminal work by Beckman et al. and Ischiropoulos et al. etc. (Ischiropoulos, H. et al., 1992; Beckmann, J.S. et al., 1994; Kers, J.A. et al., 2004) established the concept that tyrosine nitration could be promoted in vivo, which was also suggested in a previous work (Ohshima, H. et al., 1992). Although the molecular mechanism of nitration and denitration are still questioned and discussed (Pfeiffer, S. et al., 2000; Thomas, D.D. et al., 2002; Radi, R. et al., 2004; van der Vliet, A. et al., 2006; Gunaydin, H. et al., 2009; Abello, N. et al., 2009), it is convinced that protein tyrosine nitration play critical roles in physiology and pathology (Radi, R. et al., 2004; Abello, N. et al., 2009; Ischiropoulos, H. et al., 2009; Reynolds, M.R. et al., 2007; Lu, N.H. et al., 2007). Many studies had attributed nitration to loss of function (Xu, S. et al., 2006; MacMillan-Crow, L.A. et al., 1996; Eiserich, J.P. et al., 1999), but the concept as "gain of function" through tyrosine nitration was also raised through some reports (Vadseth, C. et al., 2004; Hodara, R. et al., 2004; Souza, J.M. et al., 2000). Though a relatively limited number of protein and a few tyrosine could be nitrated, this special post-translational modification has been assigned a lot of biological roles from being used as a biomarker oxidative stress to altering the structure and function, as well with influencing tyrosine phosphorylation. Current progresses show that protein tyrosine play important roles in species of biological process, including immunomodulation (Nagaraj, S. et al., 2007), apoptosis (Yakovlev, V.A. et al., 1992), cell cycle (Kong, S.K. et al., 1996), cell death (Liu, B. et al., 2009), and aging (Drew, B. et al., 2002; Hong, S.J. et al., 2007).
In this work, we manually collected 1066 experimentally indentified protein nitration sites in 554 unique proteins from scientific literature. A previously self-developed GPS (Group-based Prediction System) algorithm was employed with great improvement. We calculated the leave-one-out validation and 4-, 6-, 8-, 10-fold cross-validations to evaluate the prediction performance and system robustness. The leave-one-out validation result is accuracy (Ac) of 76.51%, sensitivity (Sn) of 50.09%, and specificity (Sp) of 80.18%. The online service and stand-alone packages of GPS-YNO2 were implemented in JAVA 1.4.2 and freely available at: http://yno2.biocuckoo.org/.
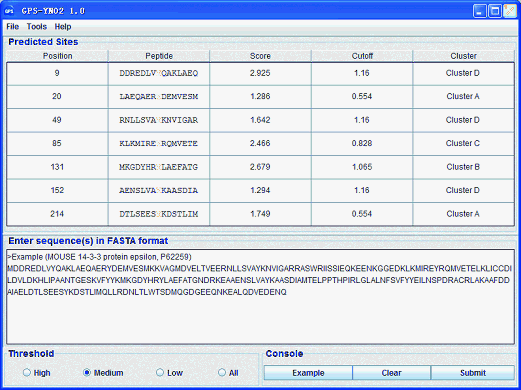
GPS-YNO2 User Interface
For publication of results please cite the following article:  GPS-YNO2: Computational prediction of tyrosine nitration sites in proteins
GPS-YNO2: Computational prediction of tyrosine nitration sites in proteinsZexian Liu, Jun Cao, Qian Ma, Xinjiao Gao, Jian Ren and Yu Xue. Molecular BioSystems, 2011, 7(4): 1197-1204. (Top ten accessed articles in August) |